June 29, 2021
Written by: Nitsan Goldstein
We all appreciate the complexity of the human brain. While our hearts, lungs, and livers are very similar to those of other mammals, our brains are what distinguish us from our primate ancestors. Humans learn, communicate, adapt, and connect with each other like no other species on Earth. But until recently, the true complexity of the human brain was still a mystery. Scientists often use animal models to study the brain because of how difficult it is to gain access to human brains both technically and ethically. If we want to study the human brain, we use non-invasive imaging to get a sense of what the brain looks like and what areas might respond to certain stimuli. Although new imaging technologies lead to major advances in knowledge, little is actually known about how individual neurons in the human brain are connected to each other and to other cells types and brain structures. This is because neurons are tiny; the cell body of a neuron is about one fifth the width of a human hair. Imaging of live human brains doesn’t come close to that kind of resolution, so how can we learn more about neuronal connections in the brain? Scientists at Harvard University and Google Research have combined advances in imaging and computational analysis methods to offer an unprecedented look into the complexities of the human brain at a nanometer scale1.
How were the images collected and analyzed?
Before diving into this study, it’s important to note that the work is published as a preprint, meaning that it has not yet undergone peer review. Experts in the field will review the paper and ensure that the research is sound and the conclusions are valid before it is published in a scientific journal. Since the findings were made available before this process took place, we can think about them as a “first draft” that may change in the coming months.
First, let’s discuss how this dataset was collected. The researchers wanted to image a piece of the human brain at very high resolution and reconstruct it using sophisticated computer programs. Pieces of human brains, as you might imagine, are not easy to get. Many brain sections that are removed surgically are diseased and would therefore not represent a typical human brain. However, neurosurgeons will occasionally remove a piece of healthy human cortex (the outer layer of the brain) in order to gain access to deeper structures when operating on patients with drug-resistant epilepsy. The team of researchers were able to get a cubic millimeter of healthy human cortex and rapidly preserve it. They then treated it with heavy metals, which is important for later imaging. Finally, the brain section was embedded in resin. The block of resin was then cut by a diamond knife into over 5,000 extremely thin sections and mounted on a long strip of tape that was wrapped around a reel, like old film.
The sections were then imaged using an electron microscope. An electron microscope sends a beam of electrons onto a sample. The electrons then scatter off the sample, and the pattern of this scatter is what creates the image. Using electrons instead of light to create an image dramatically increases the resolution that is possible. Electron microscopy can clearly show very small organelles like mitochondria inside cells. Importantly, electron microscopy is extremely useful for imaging synapses, the connections between neurons. Even the tiny vesicles containing neurotransmitters that travel across synapses are visible through an electron microscope. It is also possible to determine whether a synapse is excitatory, meaning that one neuron will be activated by another, or inhibitory, meaning that one neuron will be silenced by another. Electron microscopy, however, is not a fast process, and the team had over 5,000 brain sections to image. To speed things up, the group used a special microscope that simultaneously sent 61 beams to the sample instead of one, which significantly increased the area that can be imaged at once. This allowed the microscope to image up to 190 million pixels per second. Even with the extra beams and fast imaging, acquiring images of all the sections took a total of 326 days! All those thousands of highly detailed images took up about 2.1 petabytes of storage. To store that amount of data on the type of laptop I am using to write this article, I would need about 33,000 of them.
The group now had this enormous amount of imaging data, so how did they analyze it? The goal of their analysis was to align the 2D images in a 3-dimensional stack, basically putting the pictures of the thin brain slices back together in order, and then digitally reconstruct the neurons, other cells, and blood vessels that were in the piece of brain. The level of detail in the images allowed them to precisely map the synapses at each neuron across the cubic millimeter of the cortex. This was done using mostly automated computer programs that could track an individual cell through the various images, and then create a 3D reconstruction of that cell.
What did they find?
After reconstructing the entire section of brain that was collected, the team examined the types of cells they found and how they were connected to each other. Though there is far too much information to describe here, let’s look at some of the more surprising findings. One important conclusion was that the algorithms identified twice as many glial cells as neurons in this segment of the brain. Glial cells are cells in the brain that are not neurons and do not send electrical signals to other cells. They are, however, very important to normal brain function as regulators of neurons and neural transmission. This study highlights the important of studying glial cells and how they might contribute to normal and abnormal brain function.
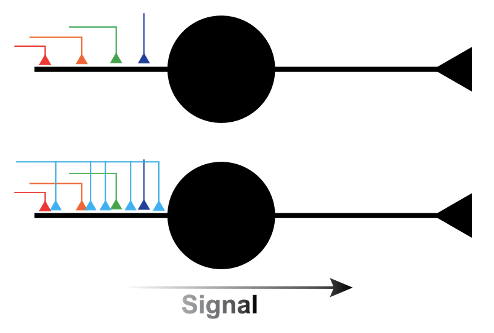
A second major finding was the sheer density of connections between neurons that were found in the sample. A total of 133.7 million synapses were identified in this cubic millimeter of human brain. Large, excitatory cells in the cortex called pyramidal neurons each received thousands of both excitatory and inhibitory inputs from the axons of other neurons. Almost all axons only formed one synapse with target neurons. Since each synapse has a relatively weak ability to change the activity of the neuron, the signal that is transmitted to the target neuron depends on the combination of these thousands of inputs. However, the group found a few exceptions where a single axon forms several (in one case up to 19!) synapses with a target neuron. This means that some inputs have a much stronger effect on the activity of the target neuron than all the other single-synapse axons (Figure 1). Though multi-synapse inputs were very rare, 30% of neurons studied had at least one input that formed 7 or more synapses. This suggests that even though these multi-synapse inputs were uncommon, many neurons could have at least one input that is significantly stronger than all the others. Discoveries like these can only be made with this kind of technique, where the source of each synapse can be tracked in 3-dimentional space at the high resolution needed to identify individual synapses.
Explore!
All of these thousands of neurons and millions of connections between them that took almost a year to image and petabytes to store came from a single cubic millimeter of a 45-year-old woman’s brain. An adult brain is about 1200 cubic centimeters, or 1.2 million times the volume of brain that was imaged in this study. It is impossible to imagine (with our brains, I might add) the amount of computation that happens inside our skulls. However, research like this at least gives us an idea of the kind of complexity that makes us human. And now, everyone can explore the data on their own! The website the team created allows you to visualize the neurons, glia, and blood vessels in 3D and even see the electron microscope images that generated the reconstructions. To see the synapses that communicate with a pyramidal neuron, click here. You can double click on cells in the microscope image or 3D image to make them appear or disappear as well as zoom in and out and scroll through the image in all 3 dimensions. Just make sure you don’t have anything urgent to do first!
References
- Shapson-Coe, A., et al. A connectomic study of a petascale fragment of human cerebral cortex. bioRxiv 2021.05.29.446289; doi: https://doi.org/10.1101/2021.05.29.446289
Cover photo by Gordon Johnson from Pixabay
Leave a comment